A comparison of per sample global scaling and per gene normalization methods for differential expression analysis of RNA-seq data
PLOS ONE: The hepatic transcriptome of the turkey poult (Meleagris gallopavo) is minimally altered by high inorganic dietary selenium
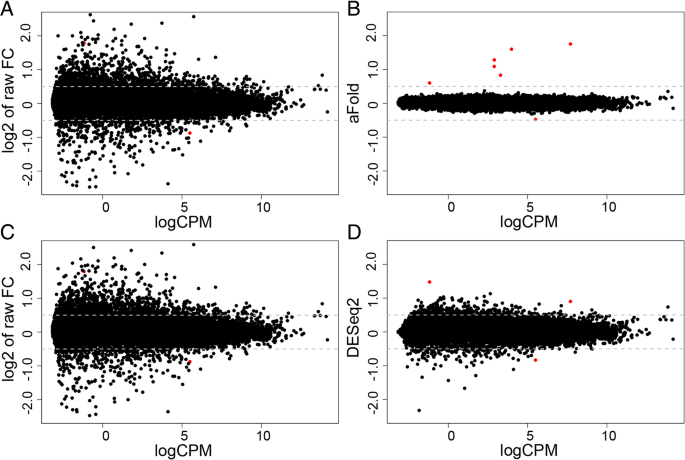
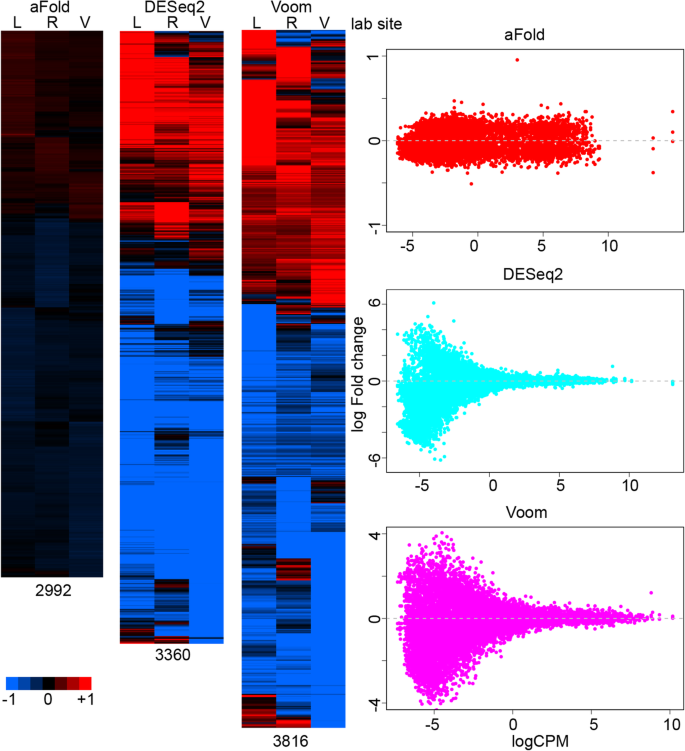
aFold – using polynomial uncertainty modelling for differential gene expression estimation from RNA sequencing data | BMC Genomics | Full Text

Differentially expressed protein groups (q < 0.4 and Log 2 fold change... | Download Scientific Diagram

Differentially expressed protein groups (q < 0.4 and Log 2 fold change... | Download Scientific Diagram

MD plot showing the log-fold change and average abundance of each gene.... | Download Scientific Diagram

Tuning Transcriptional Regulation through Signaling: A Predictive Theory of Allosteric Induction - ScienceDirect

Advantages of RNA‐seq compared to RNA microarrays for transcriptome profiling of anterior cruciate ligament tears - Rai - 2018 - Journal of Orthopaedic Research - Wiley Online Library
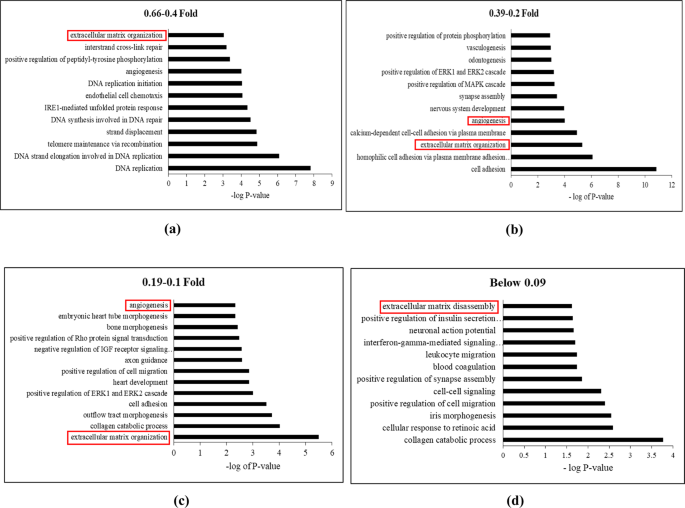

Transcriptomic Profiling of Adipose Derived Stem Cells Undergoing Osteogenesis by RNA-Seq | Scientific Reports